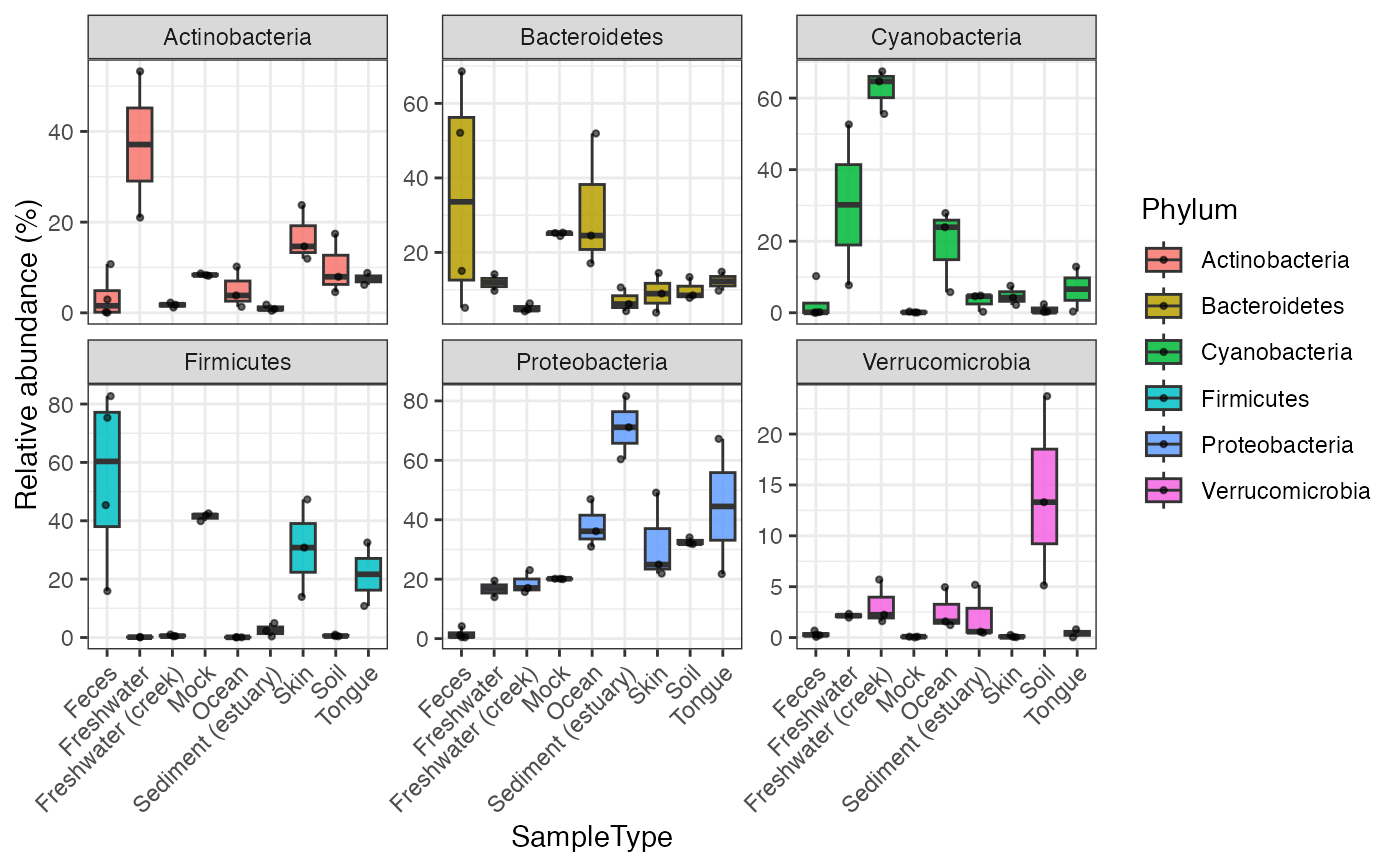
Draw sample-wise abundance distributions for selected taxa.
Arguments
- object
A
microbiome_datasetobject.- taxonomic_rank
Taxonomic rank used for aggregation.
- taxa
Optional taxa to keep. Defaults to the most abundant taxa.
- top_n
Number of taxa shown when
taxais not supplied.- x
Sample metadata column shown on the x axis.
- geom
Geometry used to display abundance.
- relative
Whether to show relative abundance.
Examples
data("global_patterns", package = "microbiomedataset")
plot_abundance(
object = global_patterns,
taxonomic_rank = "Phylum",
x = "SampleType",
top_n = 6
)
#> Ignoring unknown labels:
#> • colour : "Phylum"