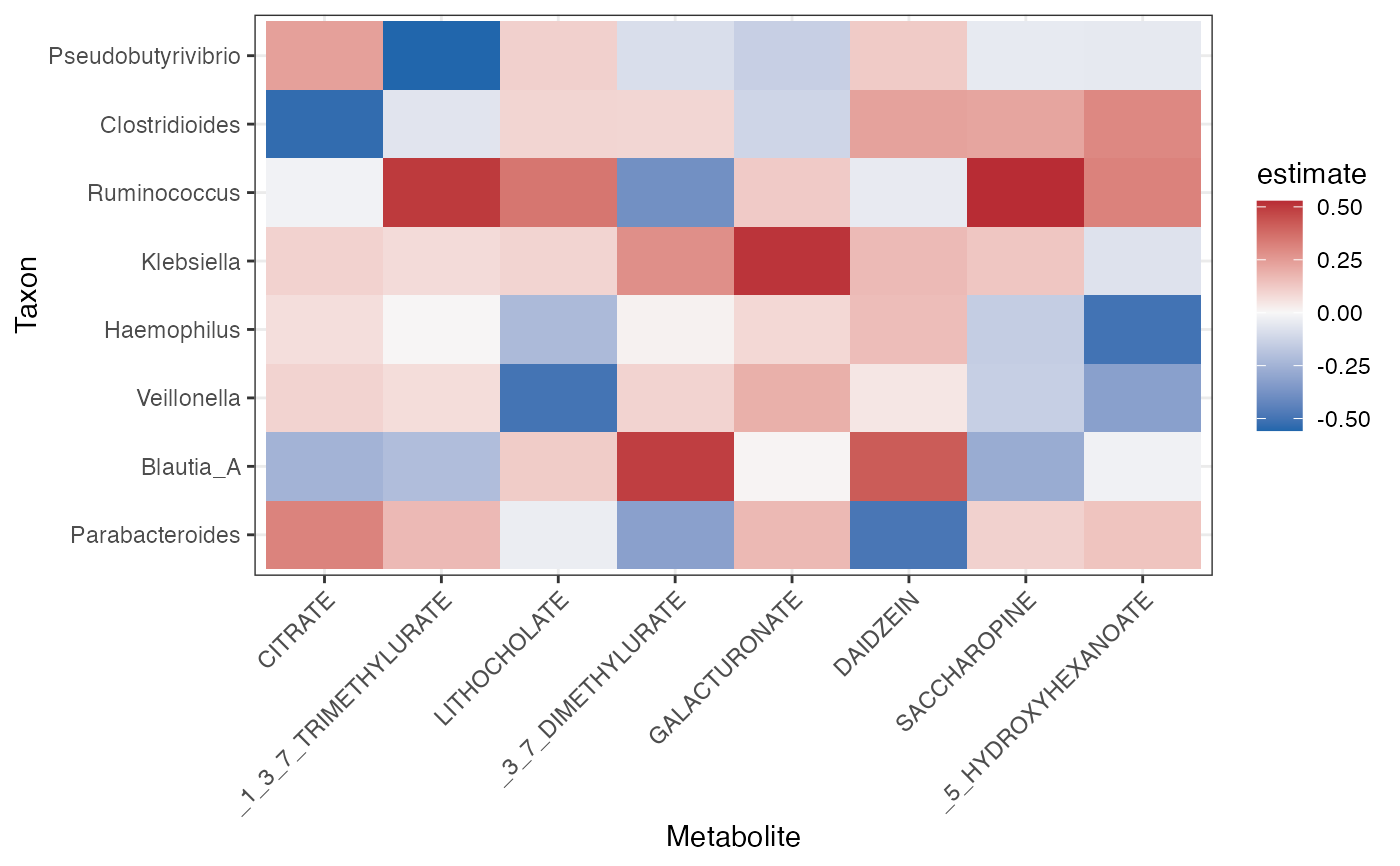
Draw a heatmap of top microbe-metabolite correlations.
Usage
plot_correlation_heatmap(
object,
top_n_taxa = 20,
top_n_metabolites = 20,
value = c("estimate", "q_value")
)Examples
data("demo_crossomics", package = "microbiomedataset")
correlation <- calculate_correlation(
microbiome_data = demo_crossomics$microbiome_data,
metabolome_data = demo_crossomics$metabolome_data,
sample_link = demo_crossomics$sample_link,
microbiome_rank = "Genus",
metabolome_transform = "none"
)
plot_correlation_heatmap(correlation, top_n_taxa = 8, top_n_metabolites = 8)