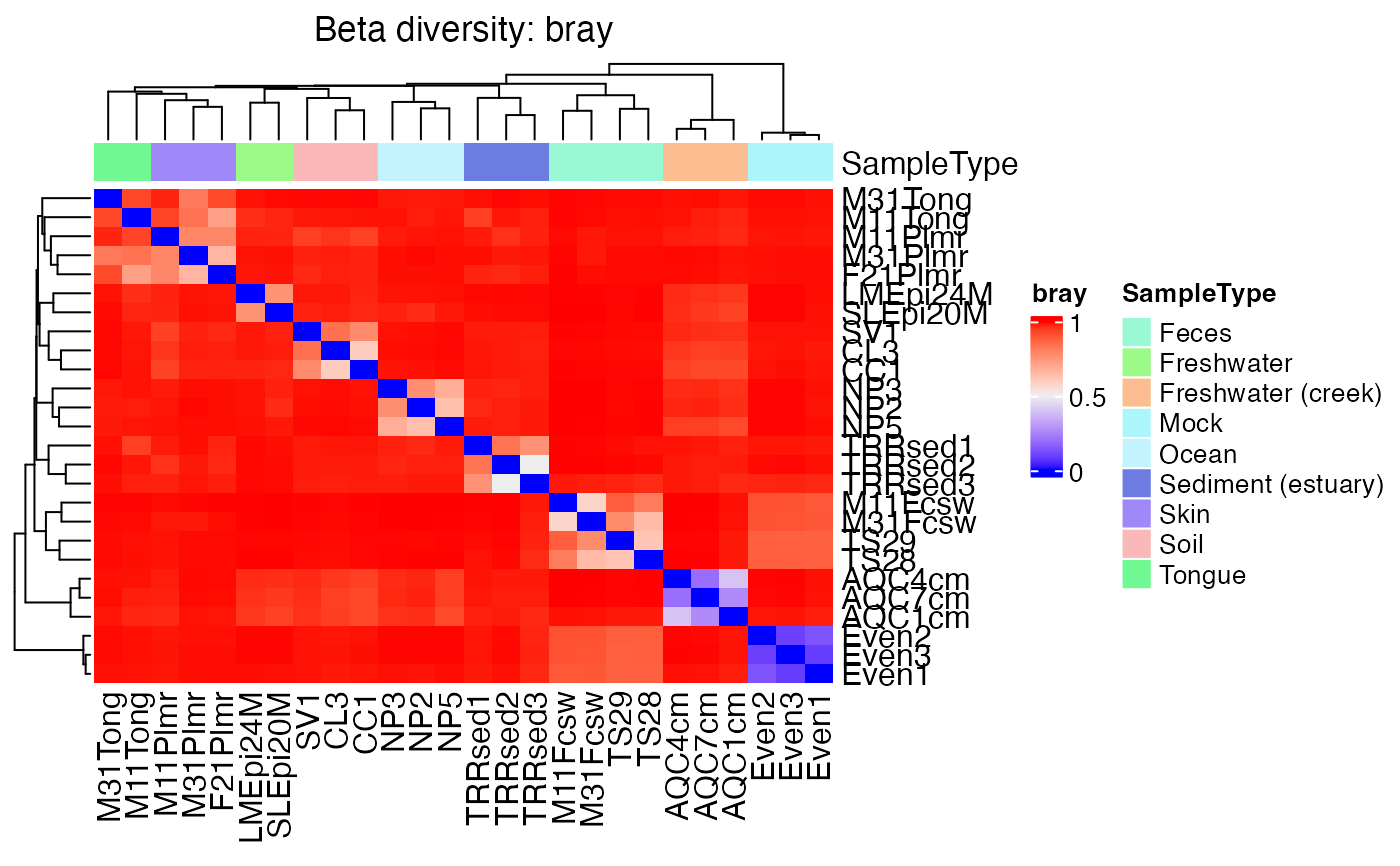
Visualize sample distances as a ComplexHeatmap heatmap.
Usage
plot_distance_heatmap(
object,
method = c("bray", "jaccard", "euclidean"),
annotate_by = NULL,
cluster = TRUE
)Value
A ComplexHeatmap::Heatmap object.
Examples
data("global_patterns", package = "microbiomedataset")
plot_distance_heatmap(global_patterns, annotate_by = "SampleType")