Show explained variance across ordination axes.
Examples
data("global_patterns", package = "microbiomedataset")
plot_scree(run_ordination(global_patterns, method = "PCA"))
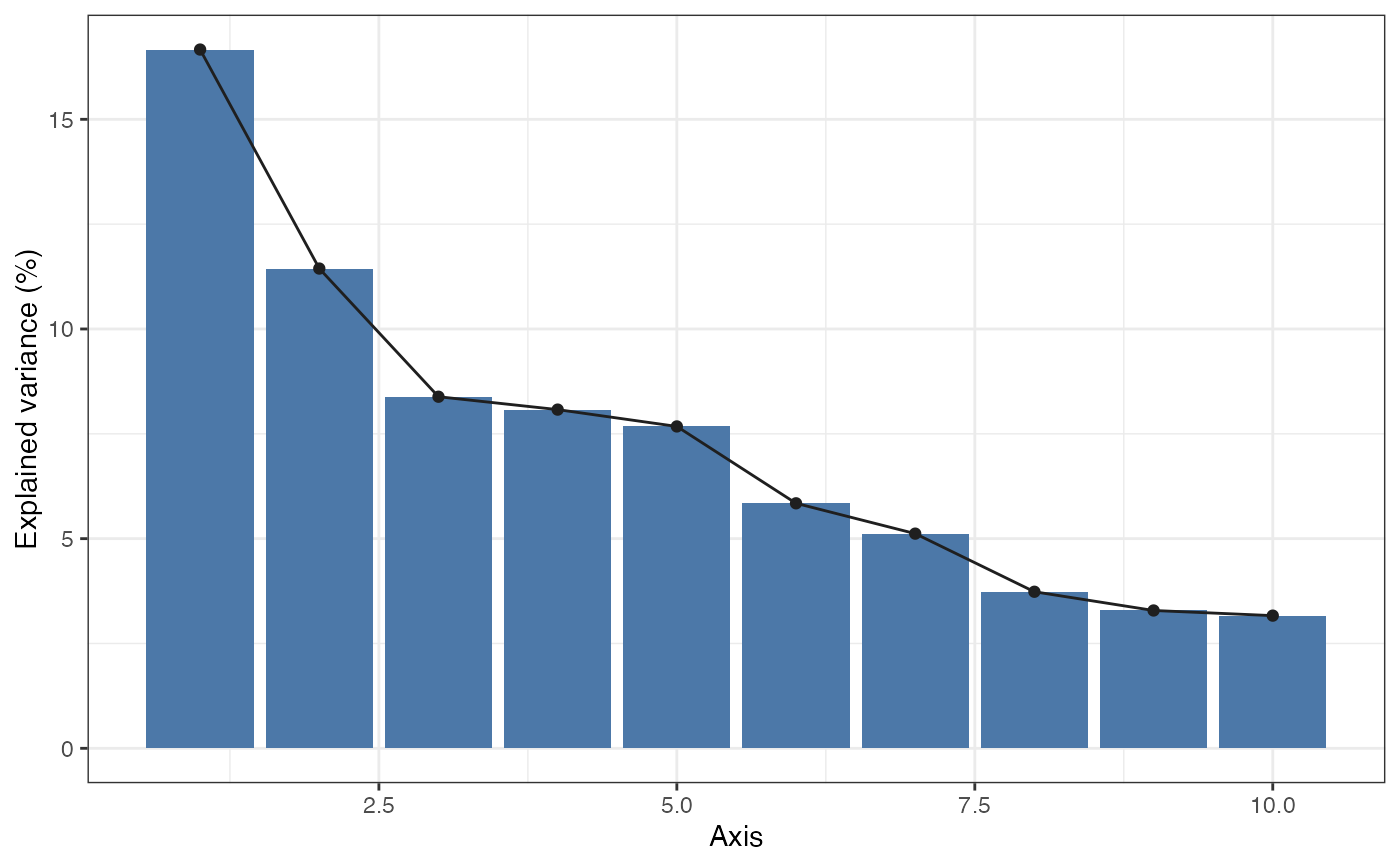
Show explained variance across ordination axes.
data("global_patterns", package = "microbiomedataset")
plot_scree(run_ordination(global_patterns, method = "PCA"))
